What do the colours on the pathway heatmaps represent? - FastBMD
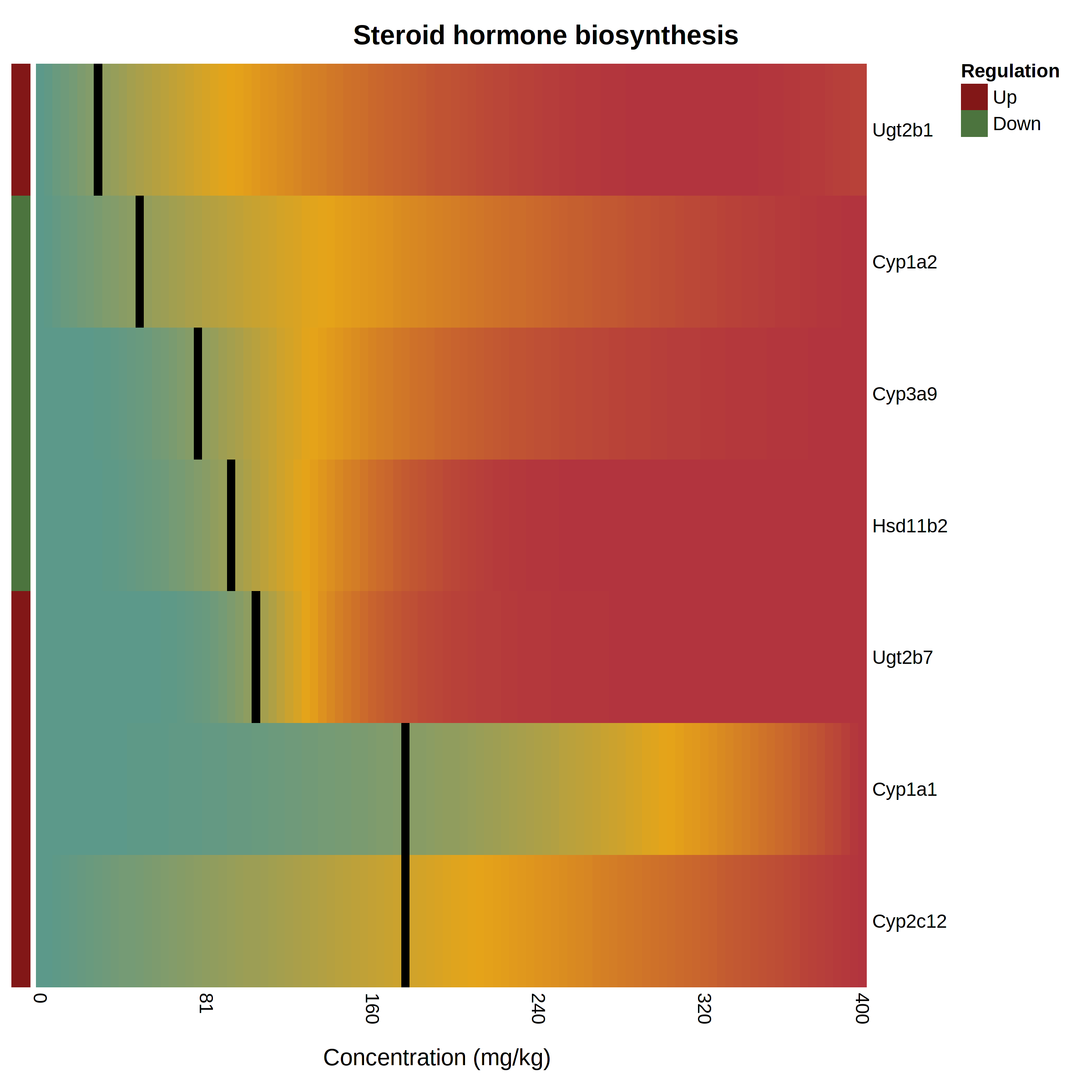
The pathway heatmap is an appealing visualization to clearly shows how the expression of each pathway gene compares to the others. It is generated when you click a pathway or gene set name in the “Gene Set Enrichment” panel at the result page. An example output is shown below The pathway heatmap values are calculated through a series of steps: The fitted model for each gene is evaluated across the range of doses in the uploaded data. The resulting modeled expression values are normalized
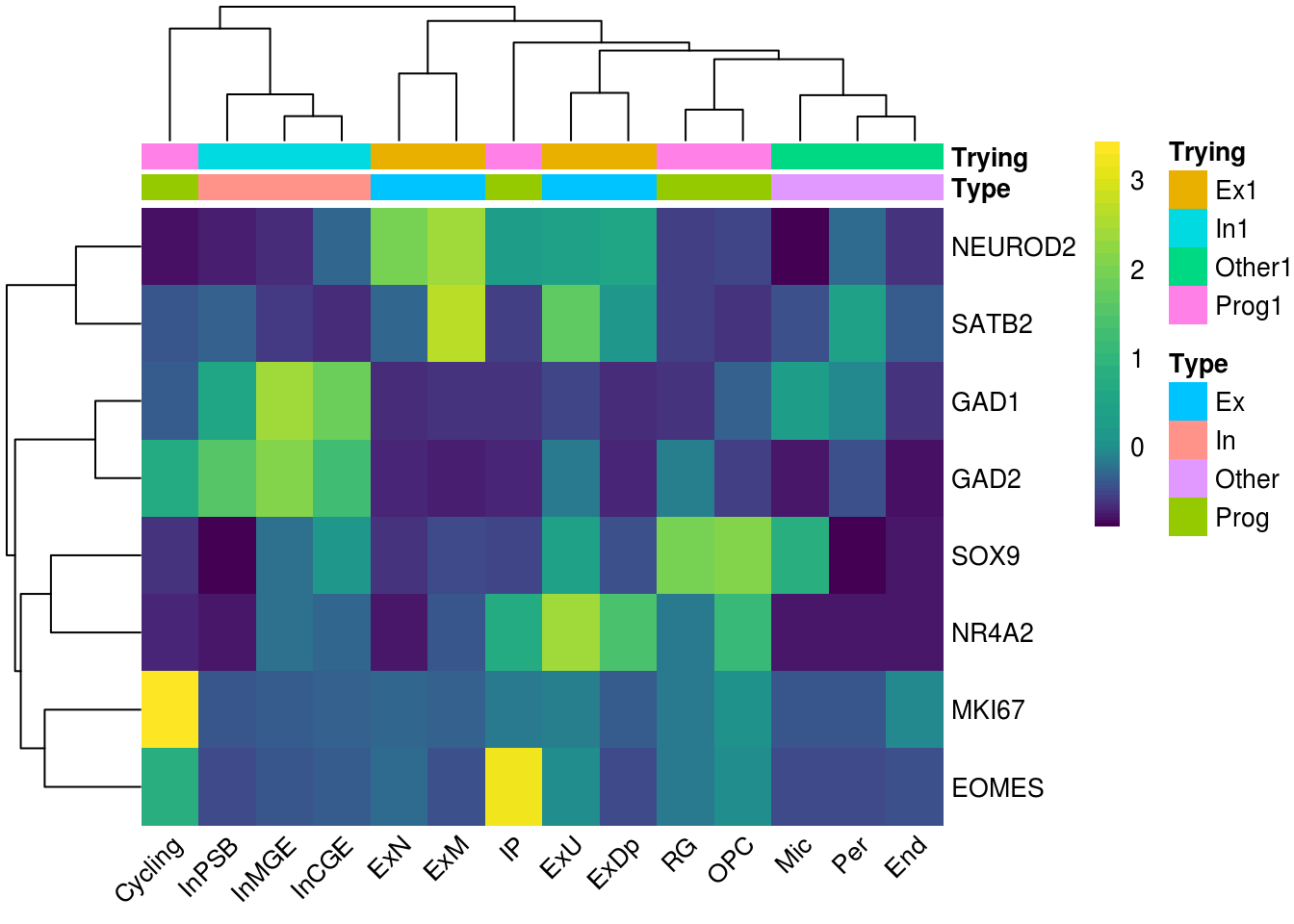
Chapter 9 Heatmap Color Palette Single Cell Multi-Omics Data

Figure S5 -Heatmap of functional categories that are

Nextcast: A software suite to analyse and model toxicogenomics
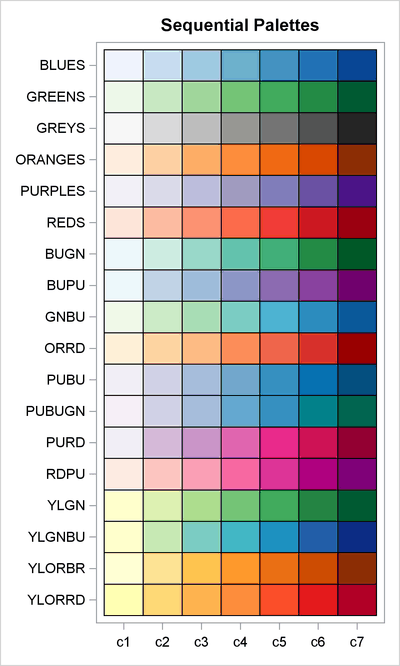

How to choose colors for maps and heat maps - The DO Loop

Latest FastBMD topics - OmicsForum

Nextcast: A software suite to analyse and model toxicogenomics
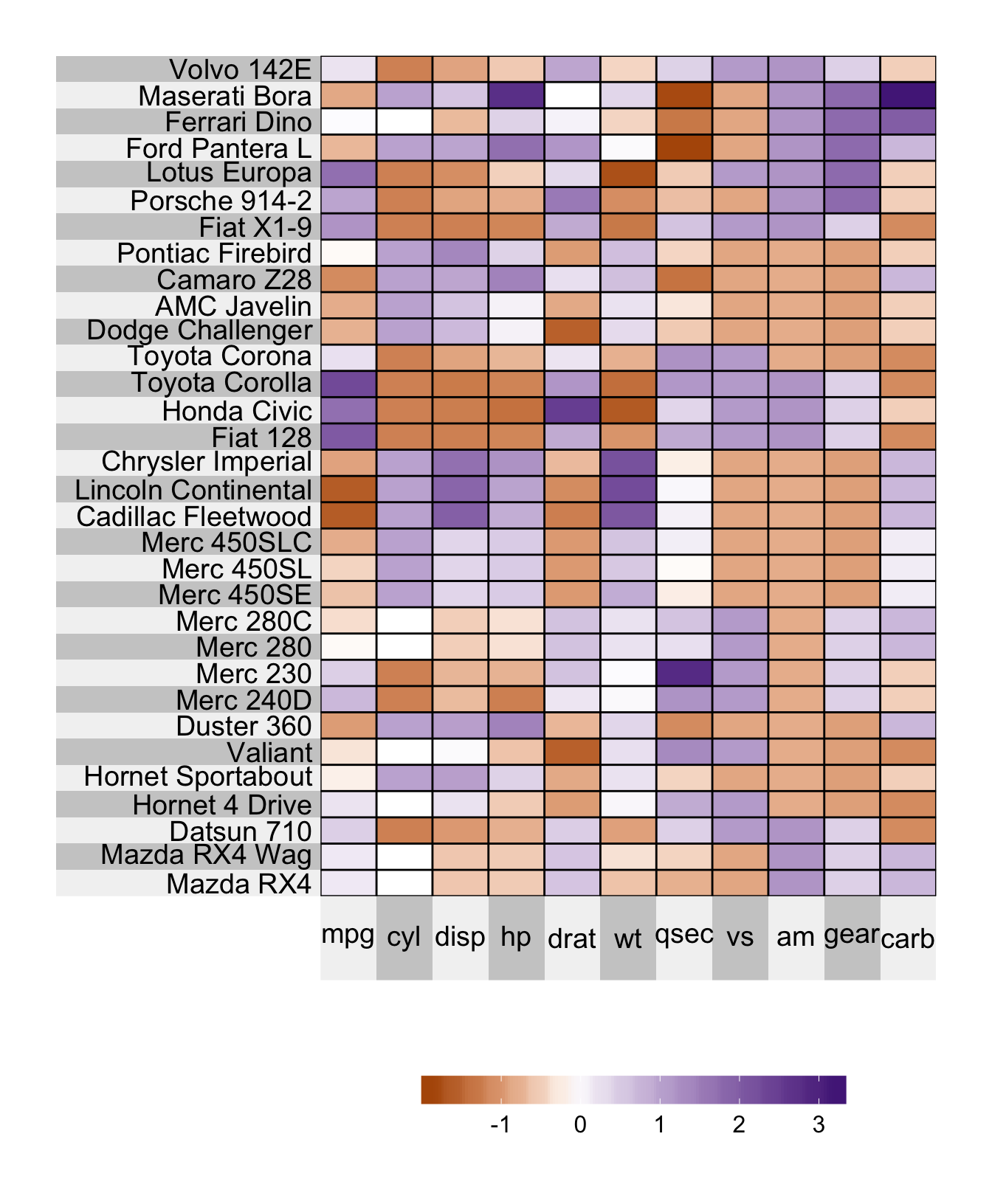
Superheat Vignette
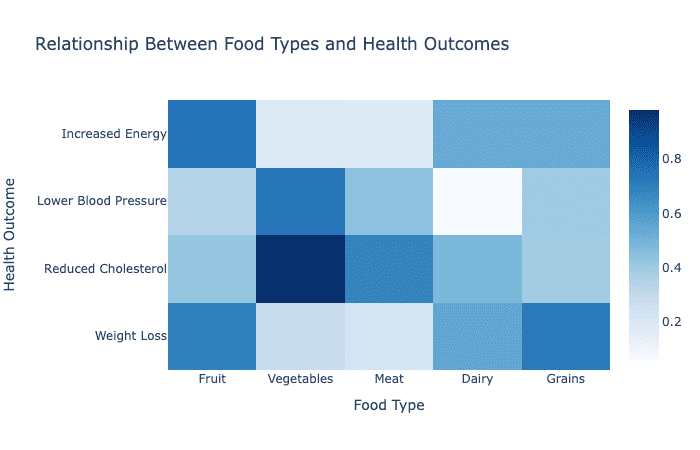
How to Use a Heatmap to Explore Data

Heat map of cluster analysis of mouse serum metabolites and
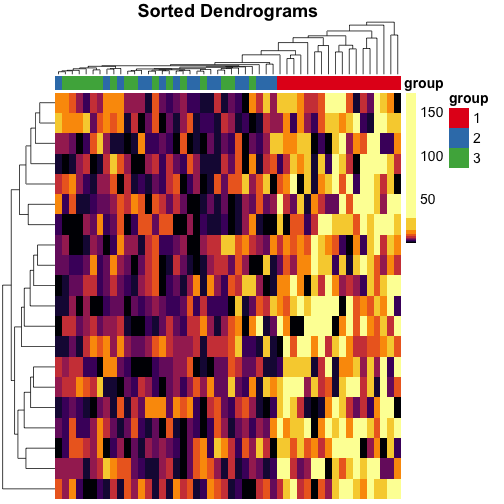
Make heatmaps in R with pheatmap - Kamil Slowikowski
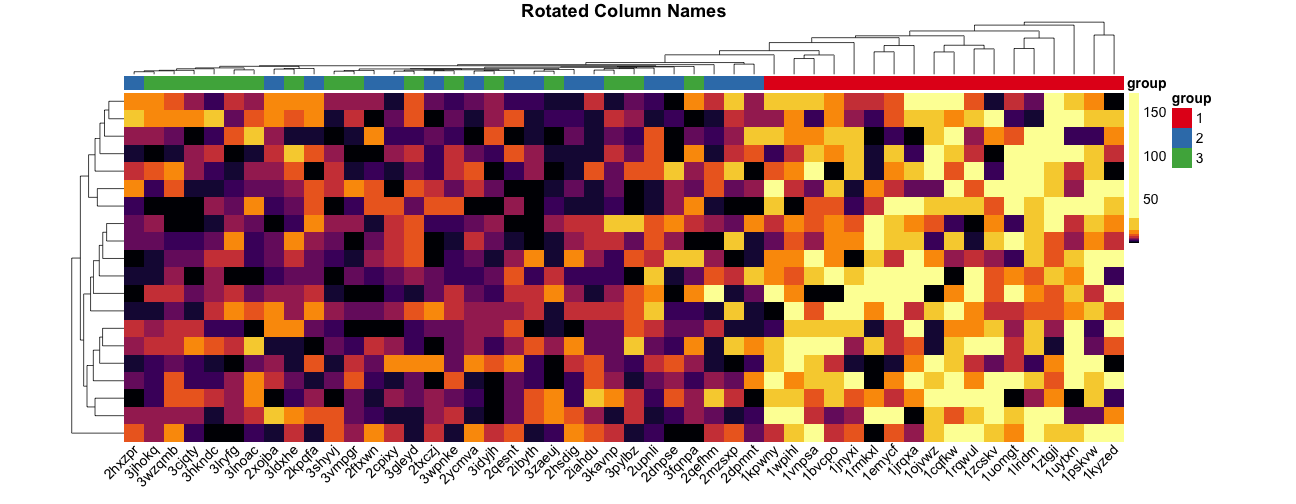
Make heatmaps in R with pheatmap - Kamil Slowikowski

Learn How to Choose Right Heatmap Colors Palette

Assessing the Toxicity of 17α-Ethinylestradiol in Rainbow Trout
Analytical Color Scales for Heat Maps - Clearly and Simply

Nextcast: A software suite to analyse and model toxicogenomics











